ECG-Federated-Learning is a Python-based open-source project with 306 GitHub stars, developed under the TTEH Lab at Dayananda Sagar University's Department of Computer Science and Engineering (Cyber Security). It tackles privacy issues in ECG-based cardiac diagnosis by implementing federated learning, where multiple simulated hospitals train models on local data without sharing raw patient signals. A global model aggregates these local updates via the Flower framework, while SHAP provides explanations for predictions to build trust in clinical use.
The approach targets IoT-enabled smart hospitals, where ECG data sensitivity under regulations like HIPAA prevents centralized training. Traditional methods risk breaches by pooling data, but this system keeps training decentralized, evaluates against centralized baselines, and maintains accuracy for detecting abnormalities like arrhythmias.
Core Features
This project combines federated learning with explainable AI for ECG classification. Key elements include:
- Federated learning pipeline using Flower, simulating multiple clients (hospitals) that train locally and send model updates to a central server.
- SHAP integration for model interpretability, generating visualizations of feature importance in ECG predictions.
- PyTorch support for high-fidelity ML models, tested in Python 3.11.
- Dual evaluation in centralized and federated modes to compare privacy-preserving performance.
- MIT-licensed code with badges highlighting Python, PyTorch, federated learning, and explainability.
These features address core needs in healthcare AI: data privacy, model transparency, and diagnostic reliability.
System Architecture
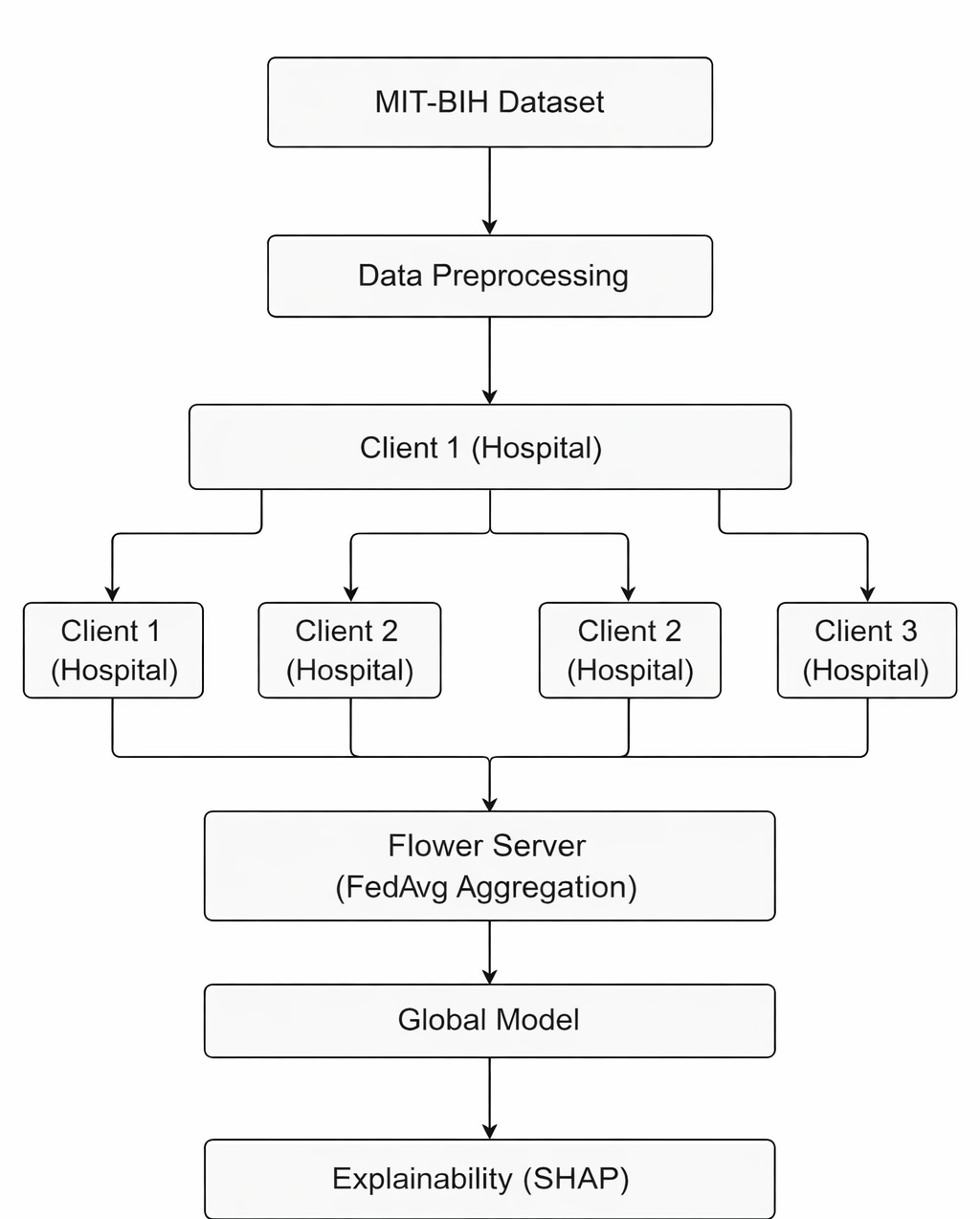
The architecture follows a distributed setup outlined in the README. Clients represent hospitals with private ECG datasets. Each runs local training on their data, using a shared model architecture. The Flower server orchestrates rounds of aggregation, typically via FedAvg (Federated Averaging).
Components include data loaders for ECG signals, client-side trainers, a central aggregator, and SHAP analyzers post-training. The README table (partially shown) maps elements like these to technologies: Flower for orchestration, PyTorch for models, and SHAP for explanations.
Simulated IoT environments mimic real smart hospitals, where devices stream ECG data without central storage. This preserves scalability for edge computing.
How It Works
Training starts with clients loading ECG datasets—public benchmarks like MIT-BIH are implied for reproducibility. Local models optimize on client data, perhaps using CNNs or RNNs suited to time-series ECG signals.
After local epochs, clients upload weight updates (not data) to the Flower server. The server averages them into a global model and redistributes it. Multiple rounds repeat until convergence.
Post-training, SHAP computes values like SHAP summary plots, showing which ECG waveform segments (e.g., QRS complex) drive predictions for classes like normal sinus rhythm or ventricular tachycardia.
The README details this in sections like "How It Works," "Core Modules," and "Code Architecture," covering data preprocessing, model definitions, and client-server scripts.
Getting It Running
The project targets Python 3.11 users familiar with ML environments. Clone the repository from https://github.com/PoorvikaN/ECG-Federated-Learning.
Setup follows the "Setup & Usage" section in the README. Create a virtual environment:
python -m venv fl_env
source fl_env/bin/activate # On Linux/Mac
# or
fl_env\Scripts\activate # On Windows
Install dependencies, primarily PyTorch, Flower (flower.readthedocs.io), SHAP, and ECG-specific libraries like NeuroKit2 or WFDB (inferred from context). A typical pip install might look like:
pip install torch torchvision torchaudio
pip install flwr[simulation]
pip install shap
pip install scikit-learn pandas numpy matplotlib
Run federated training via scripts in core modules, such as a client.py and server.py setup with Flower's simulation mode:
python -m flwr.simulation.app --run # Adjust per README examples
For explainability, invoke SHAP after model loading. Datasets download automatically or via manual placement in a data/ directory. The README's "Implementation Results" shows expected outputs like accuracy metrics and SHAP plots.
Test on modest hardware—a GPU accelerates PyTorch, but CPU suffices for simulations with few clients.
Performance Evaluation
Evaluations compare federated to centralized training. Federated setups match centralized accuracy (exact figures in README's "Performance Evaluation" and "Implementation Results"), validating privacy without performance loss.
Metrics cover precision, recall, F1 for multi-class ECG abnormalities. Plots from results likely include confusion matrices and ROC curves.
SHAP analysis reveals consistent explanations across clients, proving model robustness despite data silos.
Explainability in Action
SHAP values highlight influential time points in ECG signals. For instance, a model's decision on atrial fibrillation might emphasize irregular R-R intervals. Force plots and dependence plots aid clinicians in verifying AI outputs.
This transparency suits regulatory scrutiny, as black-box models face resistance in medicine.
Who This Is For
Researchers in federated learning, healthcare AI, or cyber security will find value here, especially students at institutions like Dayananda Sagar University replicating the work. Developers building IoT health prototypes can adapt the Flower + PyTorch stack for edge devices.
Use cases include simulating multi-hospital collaborations for rare disease detection or prototyping privacy-compliant telemedicine. Keywords like "Federated Learning," "ECG Classification," and "Privacy-Preserving Machine Learning" guide searchers to it.
It's less ideal for production without custom scaling—simulations suit proof-of-concepts, not live patient streams.
Comparisons to Alternatives
Centralized tools like standard PyTorch ECG classifiers (e.g., on Kaggle datasets) lack privacy; this adds Flower overhead but preserves data locality.
Other federated frameworks exist: TensorFlow Federated for broader ML, or OpenFL for healthcare. This project's SHAP focus differentiates it for explainability needs.
Vs. pure XAI libs like LIME, it embeds explanations in a full FL pipeline. Heavier than lightweight ECG apps like HeartPy (Python ECG processor), due to FL simulation.
Limitations noted in README include simulation-only (no real devices), potential non-IID data challenges across clients, and compute demands for many rounds.
The source code and full results reside at https://github.com/PoorvikaN/ECG-Federated-Learning, offering a solid starting point for privacy-focused ECG AI experiments.
Comments